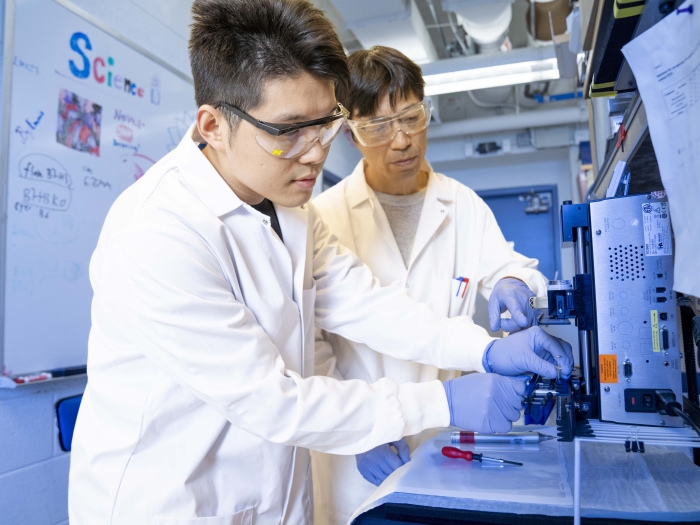
Advanced Genomics Core

Facilitating research in cutting-edge fields of genetics and genomics by implementing complex technologies.
About Us
The Advanced Genomics Core is well-known for its “next-generation” sequencing (NGS), which has revolutionized research in virtually all biological fields in the past decade. By housing NGS and companion genomic technologies in a single facility, we provide centralized expertise in advanced methods that enable all researchers to effectively exploit them to make scientific discoveries.
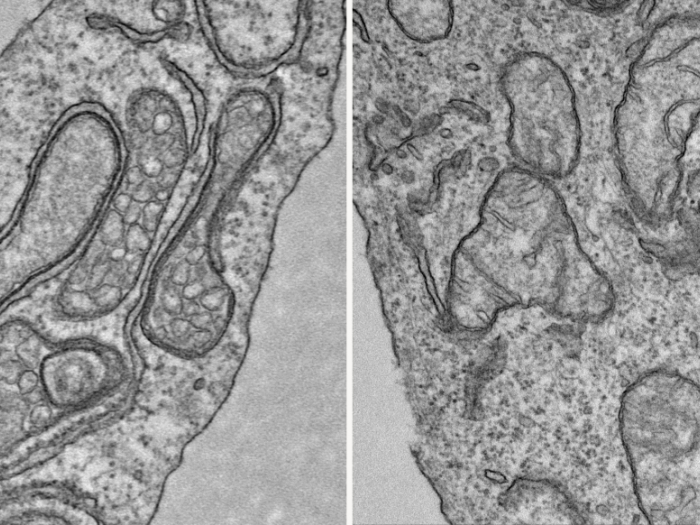
FDA-approved cancer drug fedratinib reshapes how cell organelles communicate, providing new therapeutic avenues
The anti-cancer drug alters communication between organelles that control specific metabolic pathways.
“The BRCF has been an incredible partner in our sequencing efforts from large-scale bulk sequencing to boutique full-length splice isoform studies of single-cell data sets. And all this in times of a pandemic and for studies of our patients with COVID associated AKI.”
Getting Started
How we Serve Your Research Needs

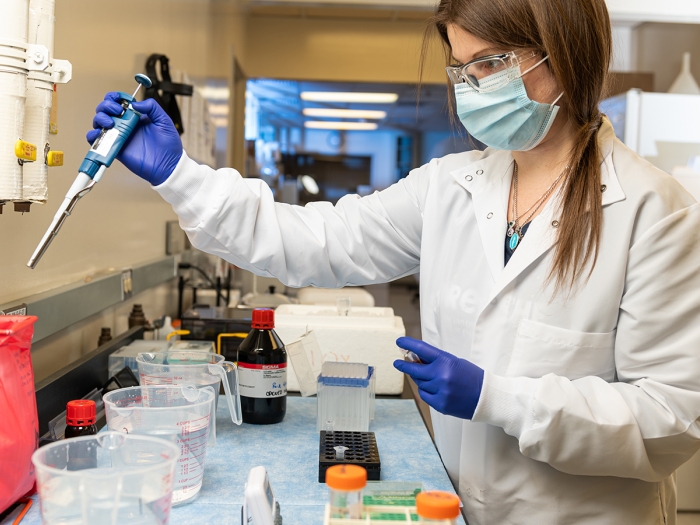
Sample Submission
General information for recommended input requirements and submission logistics.

Data Retrieval
Explore the storage and support options available when dealing with many terabytes of data.
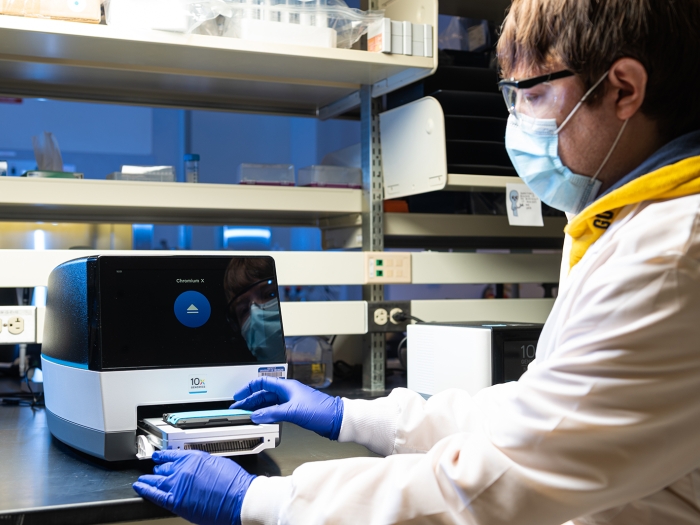

Technologies
Information about Sequencing, Single Cell, and Spatial Platforms.
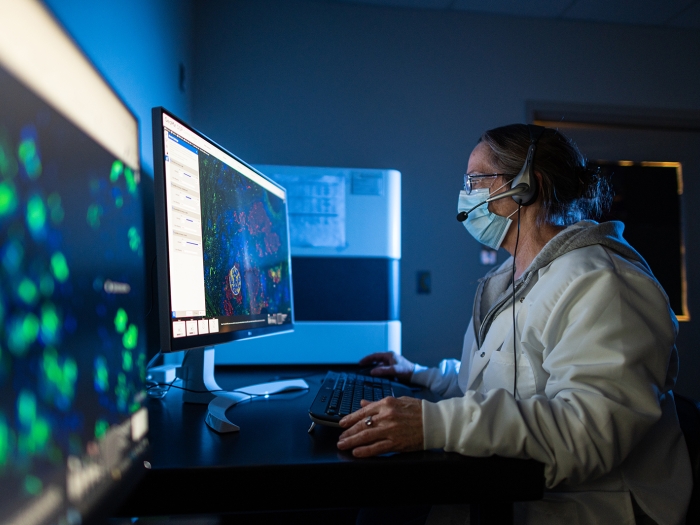

Project Estimates
Advanced Genomics Core pricing tool for estimating the cost of projects.
The AGC has been an excellent provider for our team’s sequencing needs including support for library preparations and sequencing. Because of these core services, we have been able to focus our effort on validation approaches and understanding the biological significance of sequencing-based discoveries.
Research Support & Core Policies
Letters of Support Requests
The BRCF Advanced Genomics Core is happy to provide letters of support for grant applications.
Email [email protected] to request a letter.
Please include your name and affiliation, the type of application you are submitting, the title of your proposal, the core services you will be using, and a brief description of your project.
Facilities & Resources
The Advanced Genomics Core (AGC) at the University of Michigan, a unit within the Biomedical Research Core Facilities (BRCF), provides centralized, cost-efficient access to advanced genomic technologies and expertise for investigators across the institution. Located in ~11,000 sq. ft. at the North Campus Research Complex, the AGC is a full-service facility staffed by 28 FTEs and led by Thomas Wilson (Faculty Director) and Olivia Koues (Core Director).
The core supports a wide range of applications, including bulk and single-cell RNA-seq, spatial transcriptomics, microarrays, and custom next-generation sequencing (NGS) workflows. Dedicated informatics staff perform base calling, quality control, and initial data processing, ensuring reliable and secure data delivery to users.
The AGC operates within the broader BRCF structure, one of ten integrated core units under the Medical School Office of Research, which provides centralized administrative, financial, and strategic support. This structure includes faculty-driven advisory oversight and coordinated capital investment planning, enabling continuous adoption of emerging technologies. Reflecting this, the AGC is actively expanding capacity and capabilities, including recent additions of the Ultima Genomics UG100 platform for high-throughput sequencing and advanced in situ spatial technologies. Together, this infrastructure ensures high-quality, scalable, and collaborative genomic research support aligned with institutional priorities.
Rigor & Reproducibility Statement
The Advanced Genomics Core employs multiple mechanisms to ensure rigor and reproducibility in the daily operations of the facility.
All samples are subjected to quality assessment upon entering the facility, and customers are provided with guidelines for determining if they want to proceed with downstream assays.
Core methods are well documented in the form of standard operating procedures (SOPs), and assays include appropriate positive and negative controls and contain at least one quality control step to verify the success of the method.
AGC equipment is subjected to instrumentation management plans, which include preventative maintenance as recommended by the manufacturer.
Before the release of files, data are subjected to quality control steps, which include verifying that instrumentation performance met manufacturers' specifications and initial data processing steps that ensure assay reagents worked as expected.
Budget Estimates
Next-generation sequencing projects offer many options. Our website tool is excellent for exploring project parameters and obtaining estimates.
Experimental Design & Project Consultation
The Advanced Genomics Core holds weekly office hours for project planning and consultation. Please reach out to us at [email protected] if you would like to discuss how our technologies can be applied to your area of research.
Acknowledgements
Most core services use advanced technologies that are both complex and expensive.
While we offer these in a fee-for-service format that typically does not warrant authorship, we appreciate the expertise our AGC Research Specialists bring to the University of Michigan Research Enterprise.
Please support our team by acknowledging AGC services in your publication with a sentence such as the following: “Library prep and next-generation sequencing were carried out in the BRCF Advanced Genomics Core at the University of Michigan.”
For studies that include our single cell or spatial transcriptomics services, please also add the following: Research reported in this publication was supported by the National Cancer Institute of Health under Award Number P30CA046592 by the use of the following Cancer Center Shared Resource: Single Cell and Spatial Analysis Shared Resource.
General Core Policies
- Requests for processing samples not specified at the time of the submission may require additional turnaround time and fees above initial estimates.
- The core does not return unused samples or portions of samples. Only submit aliquots to the core; do not submit your entire sample.
- All samples provided by the user will be subjected to quality control. In the case that the user elects to proceed with samples not meeting minimum core requirements (i.e., fail the quality check), the core is not responsible for the quality of the preps or data originating from those samples.
- The user is responsible for downloading all data files provided by the core. The AGC maintains a copy of all data generated for 180 days (~ 6 months). While the AGC may be able to replace lost files after 180 days, it is not a general expectation and will result in additional charges.
- The user agrees to include/acknowledge the use of AGC services in any publications that result from this work.
- The user complies with all institutional, state, and federal rules governing the performance of research involving human or animal samples.
BRCF’s Advanced Genomics Core made a huge impact on our research when we were at Michigan. This is something that we miss greatly as we moved to the University of Texas-Houston last summer. The BRCF Core is a tremendous asset of the Michigan research community, and you do amazing work!
Questions?
Contact Us
University of Michigan
2800 Plymouth Rd.
Ann Arbor, MI 48109-2800
About Us
Upcoming Research Events
FFMI Industry-Ready Webinar Series: The U-M Ecosystem for Industry Partnerships

Bio-Rad Droplet Digital PCR Lunch & Learn

The Funding Forum

Principles of Spectral Flow Cytometry Lunch and Learn

FFMI Industry-Ready Webinar Series: Agreement Types and When to Use Them

The Funding Forum

FFMI Industry-Ready Webinar Series: IP, Data, and Publications in Industry-Sponsored Research

Oncology Drug Discovery & Development (3D) Workshop

The Funding Forum

Featured Research News

Chang Kim is the Next BRCF Flow Cytometry Core Faculty Director

Ten Years of Innovation Celebrated at the Frankel Cardiovascular Center
